Epidemiological data is needed to understand the status of the virus outbreak, both at the country level and globally. The epidemiological data provides insight into whether virus control measures are adequate or whether further adjustments are needed. The data helps to inform governmental policy such as when travel restrictions can be lifted.
A COVID-19 diagnosis is confirmed by the detection of the SARS-CoV-2 virus. This information is usually recorded in the patient’s electronic medical record. Total tests administered, SARS-CoV-2 positive tests, COVID-19 recoveries, and COVID-19 deaths are recorded by government agencies.
SARS-CoV-2 is excreted in feces. The quantification of SARS-CoV-2 in wastewater could create the ability to monitor the prevalence of COVID-19 in the population of an area that feeds the treatment plant using the technique of wastewater-based epidemiology (WBE).
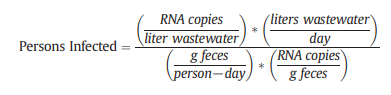
Further quantifications, molecular assay validations, and data collections for enveloped viruses in feces and wastewater are needed for accurate automated surveillance in the future. This process is driven by civil engineers, bioengineers, and computation/ data experts.
COVID-19 is a new disease never before encountered. Information is recorded in medical records. Medical doctors also write-up case reports which they publish in medical journals. Over time larger cohorts of patients are analyzed either from the medical records or under specific treatment conditions in clinical trials. These form the basis of national COVID-19 treatment guidelines, which evolve with the data released in the reports.
Data generation in medicine and biology requires the collection of samples, which need to be labeled and barcoded. Bytescout offers solutions such as Barcode Generator SDK, Barcode Reader SDK, and QR code SDK.
A diagnosis of COVID-19 is confirmed by imaging, usually computed tomography (x-rays) or ultrasound. Raw medical images are stored in a patient’s medical record, often electronically, for future reference, and as a resource for medical research and software/ hardware development if the patient grants permission.
Currently, the images are inspected manually by medical doctors, however, if enough images are collected in which the COVID-19 diagnosis is known, then a machine learning algorithm can predict the diagnosis. If the accuracy of the algorithm is high enough it can be implemented in clinical practice if it simplifies the diagnosis process. This represents an opportunity for software developers.
Certain groups in society are at increased risk during crisis situations. Those that are unable to protect themselves and others are at great risk such as those living on the street. Indirect and direct consequences of COVID-19 may increase the homeless death rate which should be carefully monitored.
Scientist driven data collection about the virus can be used to determine its structure, genes, and lifecycle, which can be used to develop treatments and vaccines. The types of data collected include genome sequence, x-ray diffraction, NMR spectroscopy, and other microscopic images of the virus, such as cryo-electron. The SARS-CoV-2 spike protein is of particular interest due to its importance in viral entry into cells.
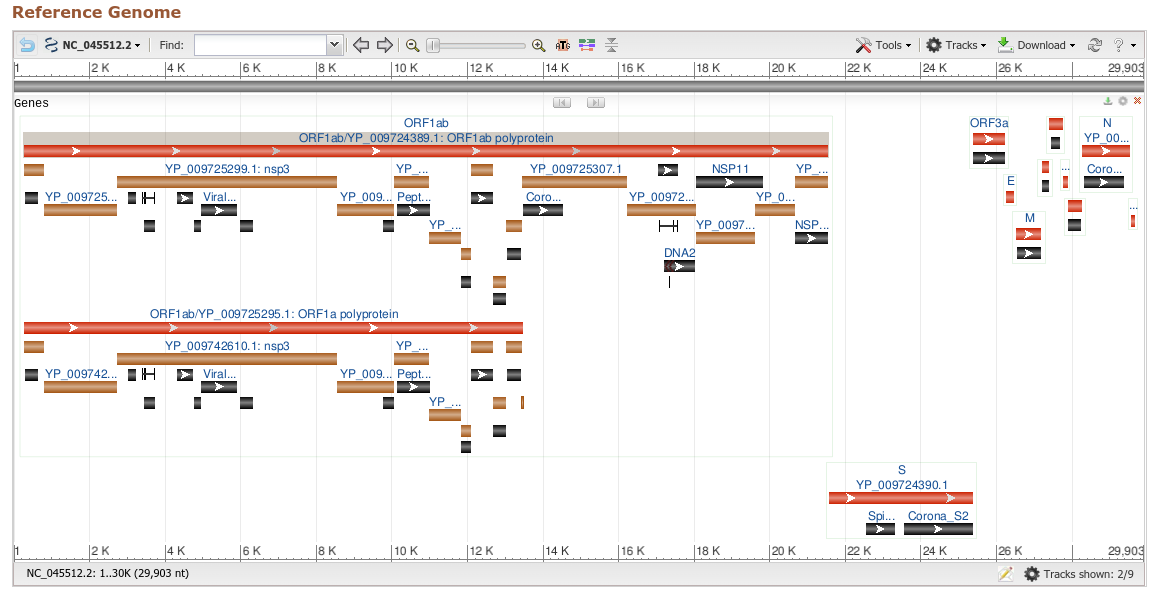
The SARS-CoV-2 genome was sequenced in January 2020. RNA viruses evolve quickly so it is important to continually sequence more virus genomes in order to monitor their evolution within new human hosts. This can provide evidence of changing viral properties. The genome can also be compared to other related virus genomes found in animals in order to determine the elusive animal reservoirs where the virus can reside and potentially re-emerge from in the future.
Cohorts of patient samples who contracted COVID-19 and their medical histories and genomic data can be correlated with outcomes. Some of these initiatives are known as biobanking. This relies on sequencing the patient in addition to the virus. Anonymized patient sequences and datasets may be submitted to GenBank. Research output is published in scientific and medical journals.
Infection data collection is largely a government, university, and hospital-level initiative, however, any organization that models risk, such as the banking and insurance sectors are likely to use the data. The availability of diagnostic test statistics varies by region, however, epidemiological data is often available through the websites of the country level healthcare agencies, such as Health Canada, which provides data for download in comma-separated values (CSV) format. The US Centers for Disease Control and Prevention (CDC) also provides a COVID-19 web-based interactive data tracker service with downloadable CSV.
Other epidemiological data sources include:
In general, centralized all-cause death rates are difficult to obtain, however, the UN does maintain country-level figures up to 2019 and predictions thereafter, up to 2100. In the USA, the Death Master File (DMF) is a computer database file made available by the United States Social Security Administration for a fee to qualified organizations. It is known commercially as the Social Security Death Index (SSDI). The file contains information about persons who had Social Security numbers and whose deaths were reported to the Social Security Administration from 1962 to present.
This is largely a medical doctor driven. Electronic health records (EHR) are now standard, and various EHR vendors distribute and maintain the software. Medical records updated by MDs are confidential unless the patient grants permission (consent) for their use in research. Research papers using anonymized data are published daily in the medical journals. Some consented EHR data can be accessed by software developers under strict controls and close collaboration with medical staff.
Socioeconomic characteristics may be noted in population-based biobank studies and may be inferred from medical records. This information is strictly confidential unless the patient consents to participation in research.
Homeless people are often missed by conventional epidemiological surveillance. Death data for people experiencing homelessness is often limited to those who have been living in city-funded shelters – ”Shelter Deaths”. As a result, the full scope of COVID-19 among this population is unknown and is collected in sporadic initiatives. Coroners, mortuaries, and local government statistics departments are sources of information, however, it is likely that COVID-19 prevalence in the homeless population can only be estimated.
Unemployment statistics and economic outlook can be used to predict the future size of the homeless population.
The Protein Data Bank (PDB) is the single global archive for 3D macromolecular structure data
The GenBank sequence database is open access, annotated collection of all publicly available nucleotide sequences and their protein translations. It is produced and maintained by the National Center for Biotechnology Information (NCBI; a part of the National Institutes of Health in the United States). It currently contains approaching 4,000 full genome sequences of SARS-CoV-2. Also available are raw next-generation sequencing files called Sequence Read Archive (.sra), which contain SARS-CoV-2 sequences.
Using all available data sources The Center for Systems Science and Engineering (CSSE) at Johns Hopkins University, USA, provides a global COVID-19 Data Repository and web-based visualization tool. This includes a map that illustrates the number of COVID-19 cases per region, along with data on deaths and recoveries.
Currently, data from water treatment plants and regarding homelessness in times of COVID-19 are lacking.
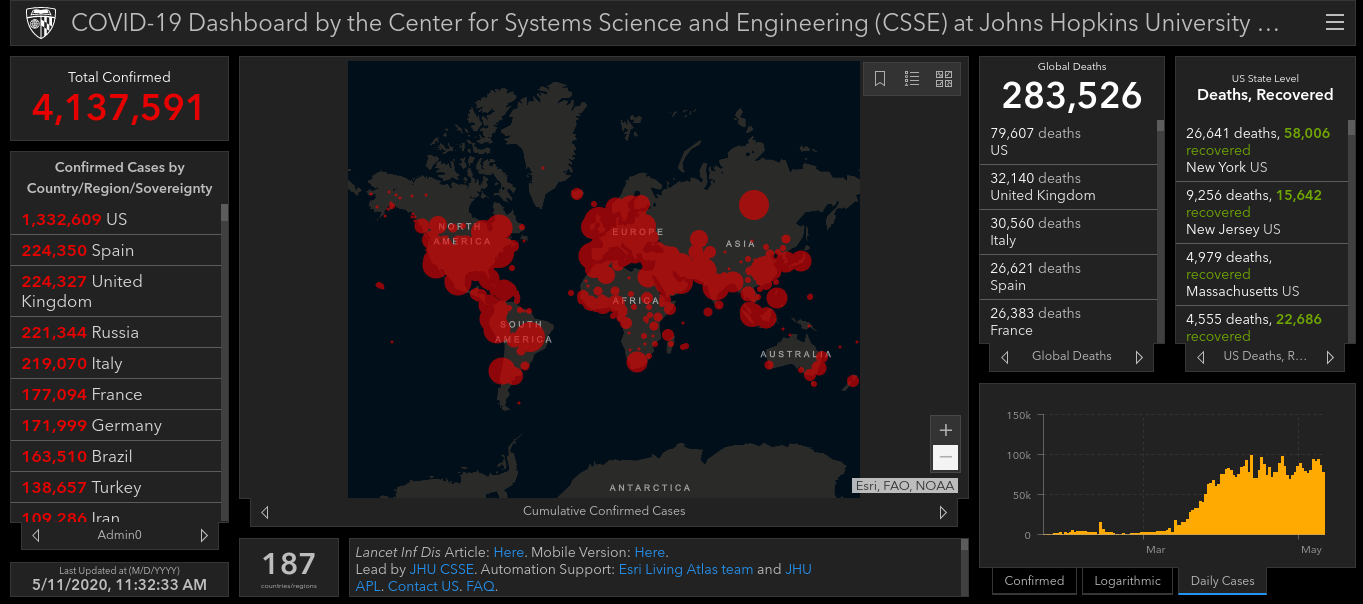
The Center for Systems Science and Engineering (CSSE) at Johns Hopkins University, USA, COVID-19 Epidemiological Dashboard
More than 300 reviewed papers on the subject of COVID-19 are typically published in scientific journals per day, which are indexed in the public library of medicine (PubMed). Pre-print servers such as bioRxiv and medRxiv host papers written by scientists but not yet officially peer-reviewed or published and are therefore potentially less reliable but crucially have the very latest information. BioRxiv currently hosts approaching 20 COVID-19 preprints per day, whereas medRxiv is currently hosting over 100 per day. At a later date, these preprints are either accepted and published in scientific journals or rejected by the reviewers.
These papers and pre-prints form the basis of the scientific rationale for the development and repurposing of therapeutics. The first wave of potential COVID-19 therapeutics largely consists of repurposed drugs and biologics due to the novelty of the SARS-CoV-2 and the need for immediate clinical action. COVID-19 is a new medical indication and as such has no currently approved therapeutics. To be approved for marketing a therapeutic must demonstrate first safety and then effectiveness in a series of three clinical trials. As of May 11, 2020, there have been over 1,358 clinical trials registered for the treatment of COVID-19. There are a number of web-hosted databases that collate registered clinical trials.
All of this data should ultimately inform the policies of the government. The major hurdle is the ability to summarize and present the front-line data, which is unavoidably complex, in a way that maintains the integrity of the information and is digestible by a non-expert. This is a fundamental challenge.
The primary index for COVID-19 data, the PubMed database, is accessible via an application programming interface (API), meaning that all of the information it contains and the linked journal articles are potentially accessible to third party programs to access natively. Normally papers are available in a “full-text” which is HTML encoded, however, given the unprecedented nature of COVID-19 many papers are only available in PDF format. In addition, the preprint servers are not indexed and all of the preprints are only available in PDF format.
Using custom software and big data techniques these sources of information can potentially be summarized in an automated and comprehensible fashion to help fight COVID-19.
ByteScout’s data extraction tools and web API are used by many research and data analysis projects in the US, Europe, and worldwide. ByteScout provides on-premise tools as well as a scalable and secure PDF.co web API to reduce the time and resources spent on manual data entry and extraction by up to 5-10x.